
Lopes, Gerardo Huck, Yue Dong, Onur Sumer and Gary D. In this demo, you can drag individual nodes, zoom in and perform other routine activities.ġ.Max Franz, Christian T. It is an improvisation over the Adobe Flash-based Cytoscape Web. The graph can be saved either in PNG or JPG format.Ĭytoscape.js is an open source software and is available at. These nodes can also be modified by removing or by selecting the interested nodes only. The graph consists of nodes representing the unit i.e., a protein or a gene. It includes the Graph theory algorithm which enables the user to search for the shortest path in the interaction graph. The graphs can also be modified by adding or deleting the graph elements such as edges. Sometimes, it is taken as an input by some core functions.įig.1 A gene-gene interaction network visualized in Cytoscape.jsĬytoscape.js offers a different type of graphs such as directed, undirected, traditional, multigraphs and hypergraphs. It allows to filter, traverse, perform various operations. Collection: It is a set of graph elements. It represents the graph, and perform various operations on the graph.Ģ. Core functions allow accessing graph elements. Core: The core is the main entry point for the developer. Many web platforms have been developed using standard technologies such as HTML, CSS, JavaScript (JS) to do this.Ĭytoscape.js is an API (Application Program Interface, i.e., it specifies how software components should interact and used to program graphical user interface), which allows the user to assimilate graphs into interaction models and web user interfaces.Ĭytoscape.js is a stand-alone tool and its architecture is broadly classified into two categories:ġ. It is easier for the researchers to study the molecular interactions in-silico, so it is equally important to visualize the network (linkage among the molecules) over modern portable devices. 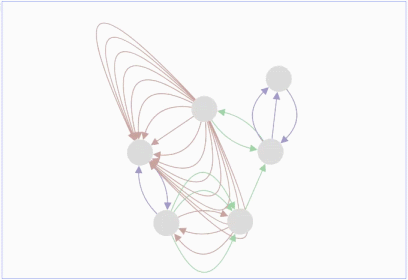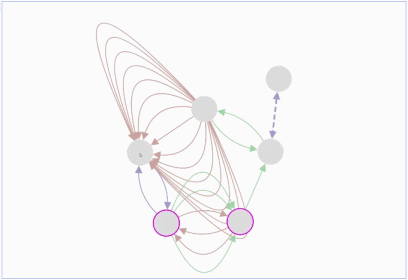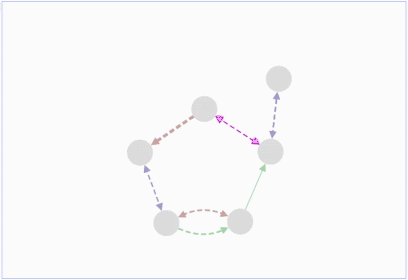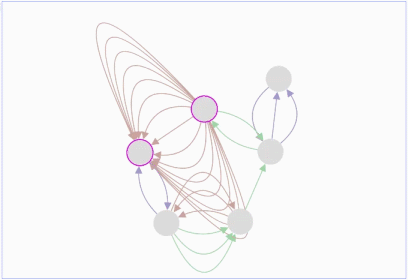
The interactive presentation used to be a dominion of Flash, It is no more same. Since the days of Flash have gone by, alternative technologies are rising. Network information is utilized in many contexts, from cellular functions to the identification of gene functions. Thus, an ever-increasing volume of the research has used network visualization to gain deep insight into the molecular interactions that influence our body functions. Network visualization has become a strong need for studying the molecular interactions whether they are the protein or gene interactions.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
